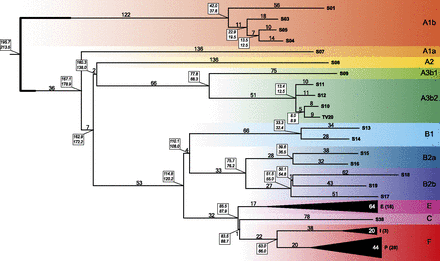
Maximum parsimony tree obtained with 2386 variable positions. The number of mutational events defining each branch is shown above it. For the collapsed haplogroups E, I, and P, the average number of mutations is shown. Dating estimates are reported in boxes near each node (upper and lower values obtained with BEAST and the rho method, respectively). Colored belts indicate major haplogroups according to current nomenclature (Karafet et al. 2008; Scozzari et al. 2012).











