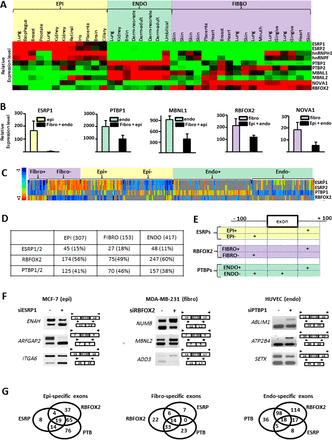
Cell type–specific expression of splicing factors. (A) Heatmap of splicing factor expression level. Each line represents a splicing factor, while each column represents a specific cell. The color of the square corresponds to the variation of the expression level of the splicing factor in each specific cell as compared to the others (green, less expressed in the cell; red, more expressed in the cell; and black, no difference). (B) RT-qPCR analysis of the expression level of ESRP1, PTBP1, MBNL1, RBFOX2, and NOVA1 in a collection of fibroblasts, epithelial and endothelial cells. (C) Spearman correlations between splicing factor expression level and the inclusion rate of epithelial-included (EPI+), epithelial-skipped (EPI–), endothelial-included (ENDO+), endothelial-skipped (ENDO–), fibroblast-included (FIBRO+), and fibroblast-skipped (FIBRO–) exons. Warm colors indicate positive correlation (e.g., a high exon inclusion level that correlates with a high splicing factor expression level), whereas cold colors indicate negative correlation (e.g., a low exon inclusion level that correlates with a high splicing factor expression level). Gray boxes indicate correlations that were discarded because of values that were not statistically significant or insufficient data available to compute correlations. (D) Summary table of epithelial-, fibroblast-, and endothelial-specific exons predicted to be regulated by the ESRP1, PTBP1, and RBFOX2 splicing factors using RNA-seq, exon array, and CLIP-seq data sets (see Supplemental Fig. S4 for more information). The number and percentage of exons predicted to be regulated by each splicing factor in each category are indicated. (E) Schematic representation of splicing factor binding site enrichment in several sets of exons differentially regulated across epithelial, endothelial, or fibroblast cells. Columns define regions in which the binding site searches were done. (F) RT-PCR analyses of the effect of depleting ESRP1, RBFOX2, or PTBP1 on alternative splicing of selected genes in the MCF-7 epithelial cell line, the MDA-MB-231 fibroblast-like cell line, or the HUVEC endothelial cell line, respectively. (G) Venn diagrams representing the number of epithelial-, fibroblast-, and endothelial-specific exons predicted to be regulated by ESRP1, PTBP1, and/or RBFOX2.











