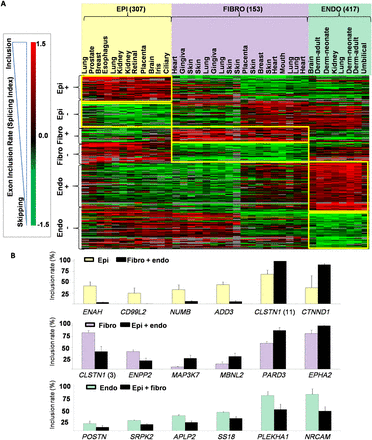
Cell type–specific splicing programs. (A) Heatmap presentation of the SI values for exons differentially spliced across fibroblast, endothelial, and epithelial cells. Exons were computationally split into several groups depending on their inclusion rate in the three major cell types. Every brace corresponds to a group (names are indicated on the left); their exons are surrounded by a yellow rectangle on the heatmap. The different categories of exons are indicated as follows: epithelial-included (EPI+), epithelial-skipped (EPI–), endothelial-included (ENDO+), endothelial-skipped (ENDO–), fibroblast-included (FIBRO+), and fibroblast-skipped (FIBRO–) exons. (B) Inclusion rate of selected exons measured after RT-PCR using cells from different origins. The gene symbol and exon position are indicated. Inclusion rates are given for epithelial cells compared to both fibroblast and endothelial cells (upper panel), fibroblasts compared to both epithelial and endothelial cells (middle panels), and endothelial cells compared to both fibroblast and epithelial cells (lower panel).











