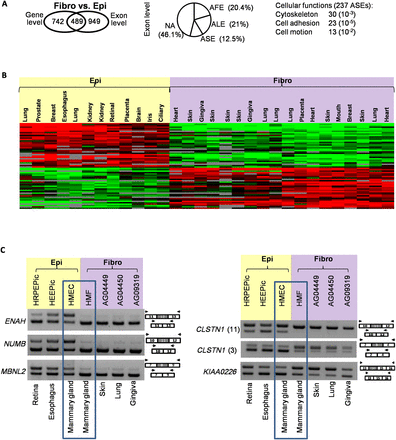
Epithelial- and fibroblast-specific splicing variants. (A) Transcriptome analysis of fibroblasts (fibro) as compared to epithelial (epi) cells at both gene and exon levels. The number of genes differentially expressed at the gene and/or exon level when comparing both cell types is shown in the left panel. The middle panel indicates the classification of events corresponding to exon level variations. Differentially expressed exons were classified according to their annotation using publicly available transcripts: alternative first exon (AFE), alternative last exon (ALE), and alternative skipped exon (ASE). Not annotated (NA) corresponds to exons that do not correspond to any of the above-mentioned categories. The cellular functions of genes differentially spliced when comparing fibroblasts to epithelial cells are indicated in the right panel. (B) Heatmap presentation of the splicing index (SI) values for exons differentially spliced when comparing fibroblast to epithelial cells. Each line corresponds to a regulated exon, while each column corresponds to a specific cell. Green boxes (–1.5 < SI < 0) correspond to a low inclusion level in the cell as compared to all the others; red boxes (0 < SI < 1.5), high inclusion level; black boxes (SI = 0), no difference for exon inclusion between cells; and gray boxes, missing values. Exons were computationally split into several groups depending on their inclusion rate that correlates with the two major cell types. (C) RT-PCR validations using RNAs from fibroblasts and epithelial cells, as indicated.











