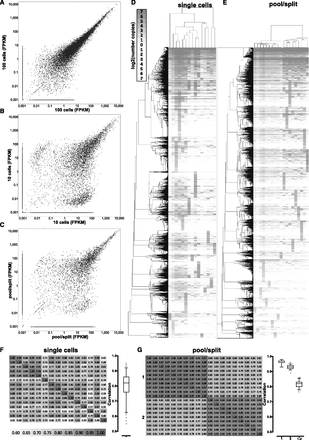
Technical and biological variation in single-cell RNA-seq measurements of gene expression. (A) Correlation between expression levels (in FPKM) between two pools of 100 cells. (B) Correlation between expression levels (in FPKM) between two pools of 10 cells. (C) Correlation between expression levels (in FPKM) between two representative pool/split libraries. A pseudocount of 0.001 was added to each data point in the scatter plots for visualization purposes. (D,E) Hierarchical clustering of estimated copies-per-cell values for protein-coding genes in single-cell (D) and pool/split (E) libraries. Pearson correlation was used as a distance metric, and only genes expressed at a level of at least one estimated copy in at least one library were included. (F,G) Correlation between estimated copies-per-cell values for protein-coding genes in single-cell libraries (F) and pool/split libraries (G). Two sets of pool/split experiments (1 and 2) are shown and “1-2” in the boxplot refers to correlations between the two sets, while “1” and “2” refer to correlation within each experiment. Similar plots, but using the Spearman correlation, are shown in Supplemental Figure 32.











