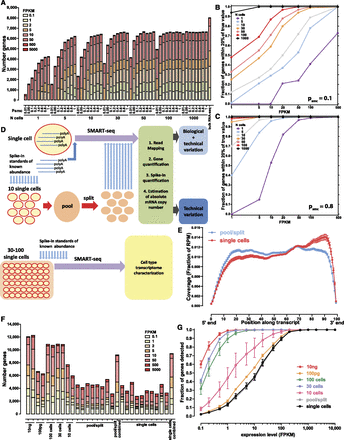
Simulated and measured transcriptome profiles from individual cells and small cell pools. (A) Number of detected genes in simulated data sets as a function of the number of cells pooled and the single molecule capture efficiency (psmc) (assuming 100,000 mRNA molecules per cell). See Supplemental Figure 1 for full details. (B,C) Accuracy of gene expression estimation as a function of the number of cells pooled and the single molecule capture efficiency; psmc = 0.1 in B and psmc = 0.8 in C, 100,000 mRNA molecules per cell assumed. Shown is the fraction of genes at the indicated expression levels in FPKM, whose estimated expression level in FPKM in simulated libraries was within 20% of their true value, after modeling the stochasticity due to the single-molecule capture efficiency of the library-building protocol. See the Methods section and Supplemental Figures 2–11 for full details. Note that the simulation is intended to illuminate the relative effects of the various parameters studied, and the absolute numbers of genes should not be directly compared to the real-life data shown in G. (D) Experimental design. Single cells are combined with spike-in quantification standards and SMART-seq libraries are generated. In parallel, multiple single cells are pooled together and combined with spikes, then lysed and split into the same number of reactions and converted into SMART-seq libraries. Libraries are then sequenced, data processed computationally, and estimates for the absolute number of copies per cell are derived based on the spikes. Variation in pool/split experiments is due to technical stochasticity, while variation in single-cell libraries is a combination of biological variation and technical noise. (E) Uniformity of transcript coverage. Shown is the average coverage along the length of an mRNA for single cells and pool/split experiments. Only mRNAs longer than 1 kb from genes with a single annotated isoform in the RefSeq annotation set were included. See Supplemental Figure 29 for more details. (F) Number of detected protein-coding genes for libraries built from 10 ng and 100 pg of poly(A) RNA, pools of 100, 30, and 10 cells, representative pool/split experiments (individually and summed across all libraries), and representative single cells (individually and summed across all libraries). (G) Fraction of genes from 100-ng bulk poly(A)+ RNA libraries that were detected in pools of 100, 30, or 10 cells, 100 pg of poly(A)+ RNA, pools/split experiments, and single cells. FPKM is shown on the x-axis.











