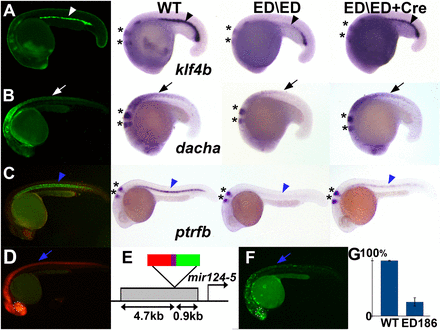
Example of insertions that cause mutations in nearby associated genes. (A) ED25 GFP expression recapitulates the expression pattern of klf4b detected by in situ hybridization, in the blood island (white and black arrowheads). (B) ED27 shows expression of GFP in the spinal cord (white arrow), which coincides with the expression of dacha (black arrow). (C) ED170 shows a strong expression of GFP in the notochord (blue arrowhead), recapitulating the expression of ptrfb (blue arrowhead). (A–C) In all three cases, homozygous embryos for the insertions show decreased levels of transcripts for the associated genes (third column), and the endogenous expression is recovered when Cre recombinase is injected in this homozygous mutant background (fourth column). Asterisks mark the expression of egr2b, a marker used as an internal control for the in situ hybridization. (D) ED186 shows strong RFP expression in the central nervous system (blue arrow) and eye (white dotted circle). (E) This line is an ED insertion near mir124-5 oriented with the RFP enhancer trap upstream of this noncoding gene. (F) In ED186, an assay for enhancer activity of the sequence 4.7 kb upstream of the insertion point reveals expression in the central nervous system (blue arrow) and eye (white dotted circle). (G) Homozygous embryos for ED186 insertion show a decrease of >70% in the transcription levels of mir124-5, detected by qPCR.











