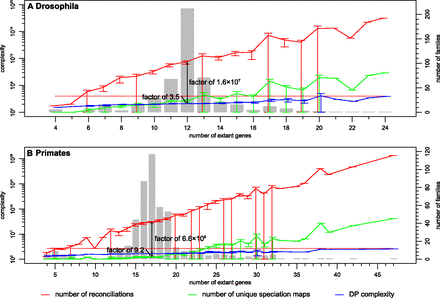
DLCpar complexity on simulated fly and primate gene trees. For each data set, gene trees were divided into classes based on the number of extant genes (counts shown as bars). For each gene tree, we determined the total number of reconciliations considered by DLCpar (red), the number of reconciliations if only relative locus maps for speciation nodes are considered (green), and the number of computations to complete the DP table (blue). The mean and 95% confidence interval (mean ± 1.96 × standard error) is shown for each class. Also shown are the number of reconciliations considered by DLCoalRecon (red dots), which was fixed at 1000 search iterations. (DLCoalRecon utilizes a prescreening approach so that 20 × 103 locus trees are proposed, but only 1000 of these are considered using the full probabilistic model.) Data are shown for flies with simulation parameters of 1× duplication-loss rate, g = 0.1 yr, N = 25 million; and for primates with simulation parameters of 1× duplication-loss rate, g = 20 yr, N = 25,000. Note that the number of reconciliations considered by DLCpar increases with increasing gene tree size (red), redundancy in the reconciliation search space is dramatically reduced by considering relative locus maps at speciation nodes (green), and dynamic programming further increases efficiency by reusing subproblems (blue).











