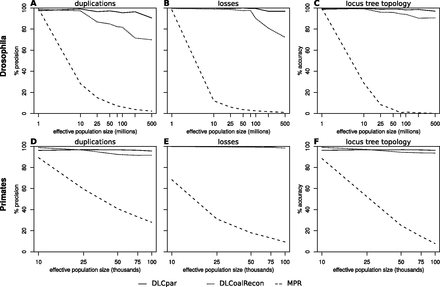
DLCpar performance on simulated fly and primate gene trees. DLCpar, DLCoalRecon, and MPR were used to reconcile simulated gene trees. Simulated data sets and DLCoalRecon and MPR results were obtained from Rasmussen and Kellis (2012), and DLCpar was run with costs of D = 1, L = 1, C = 0.5. Duplications and losses were simulated at rates estimated from real data (flies: 0.0012 events/gene/myr; primates: 0.0017 events/gene/myr), generation times for model organisms were obtained from the literature and assumed equal throughout the clade (flies: 0.1 yr; primates: 20 yr), a wide range of effective population sizes was used, and 500 gene trees were simulated per parameter setting. For the fly data set, DLCpar shows substantial improvement over DLCoalRecon in both the precision of inferring duplications and losses (A,B) as well as the accuracy of reconstructing the locus tree topology (C). For the primate data set, DLCpar and DLCoalRecon performance is comparable (D–F). Both ILS-aware methods dramatically outperform MPR.











