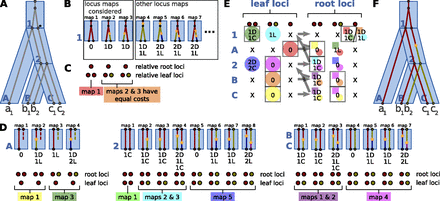
The DLCpar algorithm. In this example, we use equal costs for the evolutionary events. Furthermore, for illustrative purposes, the root of the gene tree has been extended so that a duplication may occur along the root branch; in general, this root branch is not necessary. (A) LCA mapping is used to map the gene tree (gray) within the species tree (blue), and implied speciation nodes (*) are added to gene tree branches that span multiple branches of the species tree. (B) Starting at the root branch of the species tree, DLCpar enumerates the locus maps and determines an optimal order and reconciliation cost for each. (In practice, some locus maps are not considered since they are never most parsimonious.) (C) The leaf loci are remapped to relative loci, and for each unique labeling of root loci and leaf loci, an optimal underlying locus map (and associated order) is selected. (D) This is repeated for all branches of the species tree in pre-order traversal, thereby enumerating all locus maps (along with associated optimal orders and reconciliation costs) for this gene tree and species tree. (E) For each species and each assignment of root loci and leaf loci, dynamic programming (DP) is used to determine the minimum total cost along all descendant species tree branches. The DP table is filled by post-order traversal of the species tree (arrows); see text for how these costs are computed. Colors indicate which leaf loci (circles) and which species-specific locus map (squares with colors corresponding to parts C and D) are used. At the root of the species tree, the optimal cost is selected (boxed), and traceback allows assignment of the loci for all speciation nodes (boxed). These can then be used to determine the species-specific locus maps and orders. (F) The inferred LCT is shown.











