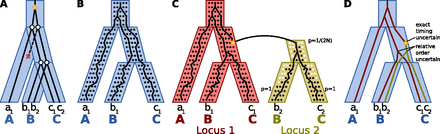
The three-tree model and the labeled coalescent tree. (A) In the duplication-loss model, incongruence between the gene tree (black) and species tree (blue) can be explained using gene duplications (yellow star) and gene losses (red “x”). (B) In a multispecies coalescent model, incongruence between the gene tree and species tree can be explained due to incomplete lineage sorting (ILS). Because no duplications or losses are allowed, this model is inapplicable to gene families in which multiple gene copies exist in at least one species. (C) The unified model proposed by Rasmussen and Kellis (2012) combines the multispecies coalescent and duplication-loss models. In this example, a duplication occurs in one chromosome [note the duplicate's frequency is initially p = 1/(2N), where N is the effective population size, assuming a diploid genome] and creates a new locus, “locus 2,” in the genome. At locus 2, the Wright-Fisher model dictates how the frequency p of the daughter duplicate (black dots) competes with the null allele (white dots) until it eventually fixates (p = 1). A gene tree is a “traceback” in this combined process. Note that the red and yellow trees form an intermediate locus tree (distinct from the gene tree and species tree) that describes how loci are created and destroyed. In this example, the gene tree has the same topology as that in A, but incongruence with the species tree is explained by duplication and deep coalescence. (D) The LCT combines the species tree, locus tree, gene tree, and reconciliations between them into a single structure. Each node of the gene tree is labeled with the species and locus to which it belongs, and gene tree nodes within the same species and locus are totally ordered in time. (Parts of this figure have been adapted with permission from Rasmussen and Kellis [2012].)











