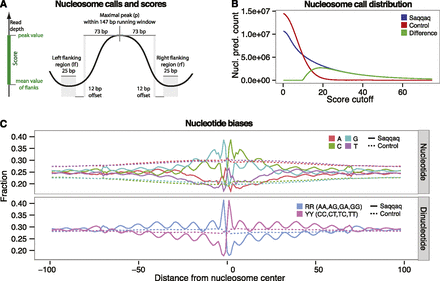
Nucleosome calls and positioning patterns. (A) Nucleosome center positions (dyads) are called as read-depth peaks if maximal at the center of a running window of nucleosome length 147 bp. Calls are scored by the difference in read depth between the peak (p) and the average read depth of the left (lf) and right (rf) flanking regions [score = p − (lf + rf)/2]. (B) Nucleosome call abundance is shown as a function of quality score cutoff for the Saqqaq (blue) and the Control (red), which lacks the nucleosome signal. The difference (green) gives the expected number of true positive calls at a given score cutoff and, indirectly, the FDR (<1% for the 1.9M calls with a score cutoff >29). (C) Base composition and distribution of purine/pyrimidine sequence dimers across the top 25% of called nucleosomes.











