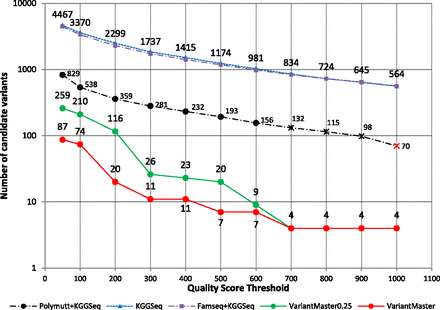
Causative recessive and X-linked variants' identification in a family presenting with a Mendelian disorder characterized by mental retardation, ataxia, and epilepsy. The number of candidate variants as a function of increasing quality score thresholds obtained by VariantMaster, KGGSeq, PolyMutt + KGGSeq, and FamSeq + KGGSeq is shown. Results obtained by VariantMaster after reduction to 25% coverage (∼10×) in the unaffected individuals are shown in green. All methods except PolyMutt + KGGSeq, which lost the SLC9A6 variant at QS > 700 (black cross) and the ADSL variant at QS > 1000 (red cross), identified the two causative variants at each threshold (for details, see text).











