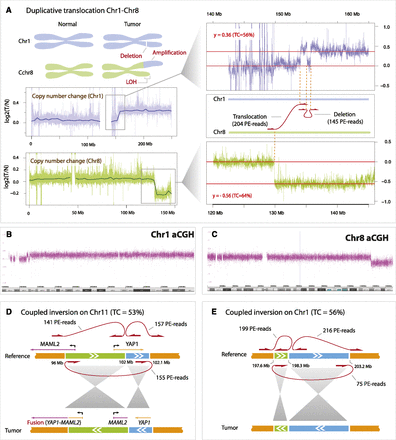
Examples of structural variants detected within NPC-5989. (A) Schematic representation of chromosomal rearrangements affecting Chr1 (blue) and Chr8 (green). Coverage plots demonstrate an increase in copy number of Chr1q and a reduction of copy number of Chr8q. Each data point represents the log ratio of tumor read counts to normal read counts in 5-kb bins across the chromosome. Red lines represent mean coverage corresponding to regions with different copy numbers. Based on the ratio levels, we estimate that tumor content is 56% (based on amplification of Chr1q) and 64% (based on deletion of Chr8q). The magnified regions contain two breakpoints representing duplicative translocation and deletion (shown as linked red arrows), supported by 204 and 145 read pairs, respectively. Coordinates of breakpoints match closely with positions of copy number changes both on Chr1 and Chr8, supporting the same rearrangement event. Deletion breakpoint coordinates also match coordinates of the region on Chr1 where coverage visibly drops. The duplicative translocation results in a region of LOH 18 Mb in size at the end of Chr8, and in the amplification of much of Chr1q. (B,C) Results of the array CGH analysis of NPC-5989 on Chr1 and Chr8. The x-axis represents genomic coordinates, whereas the y-axis represents probe saturation which is converted into copy number calls. (D) Coupled inversion involving 6-Mb and 0.1-Mb regions on Chr11, which results in the YAP1-MAML2 gene fusion product. The coupled inversion is represented by three breakpoints (linked red arrows). (E) Coupled inversion involving 0.7-Mb and 4.9-Mb regions on Chr1.











