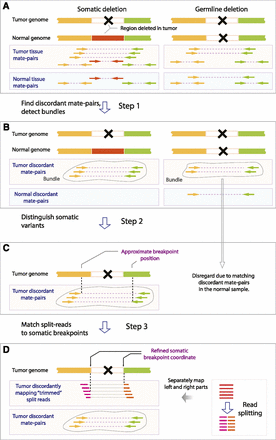
(A) SMASH workflow illustrated by comparing a hypothetical somatic structural variant (left portion of the figure) and a hypothetical germline structural variant (right portion of the figure). The region that is deleted is pointed out by a black cross. Arrows connected by dashed lines represent read pairs, in which reads of concordant pairs have the same color and those of discordant pairs have a different color. Different colors also represent different genomic regions that have been juxtaposed by a structural variant. (B) Step 1: SMASH eliminates concordant pairs and retains discordant pairs, and groups discordant read pairs from the tumor sample into bundles (contoured by gray lines) based on proximity of underlying read coordinates and consistency of orientations. (C) Step 2: Approximate coordinates of breakpoints are derived from read bundles; then each normal read pair is compared to tumor breakpoints and all breakpoints that have normal read pairs supporting them are eliminated. (D) Step 3: Sequencing reads are split and ends are mapped independently; discordant split read coordinates are used to further refine breakpoints.











