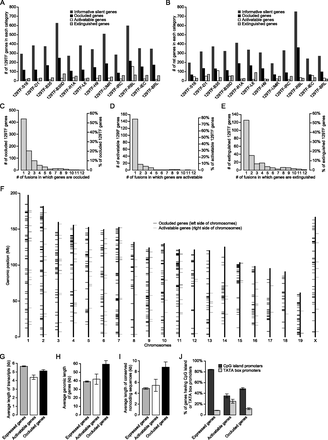
Identification of occluded genes and characterization of their structural features. (A,B) Number of informative silent, occluded, activatable, and extinguished genes in 129TF (A) or in rat cells (B) identified by each fusion. (C–E) Histograms of 129TF genes based on the number of fusions in which a given gene is occluded (C), activatable (D), or extinguished (E). (F) Genomic distribution of occluded and activatable genes in 129TF. (G–I) Average lengths of transcripts (G), average genomic lengths (H), and average lengths of conserved noncoding sequences (I) for expressed, occluded, and activatable genes. Genomic length is the distance between the annotated TSS to the annotated end of the gene body, per data provided by the UCSC Genome Browser. Noncoding sequence of a gene is the sequence from 2 kb upstream of TSS to the end of the gene body, minus all coding regions. (J) Percent of expressed, occluded, and activatable genes that possess either CpG island promoters or TATA box promoters.











