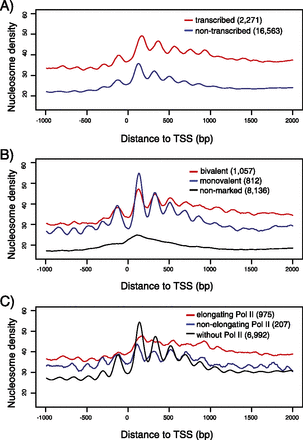
Canonical nucleosome organization at promoters in the absence of transcriptional elongation. (A) Average promoter nucleosome organization for transcribed (red) and nontranscribed (blue) genes. Transcription status is inferred from H3K36me3 ChIP-seq enrichment in exons and RNA Pol II ChIP-seq enrichment in gene bodies. (B) Average promoter nucleosome organization for three types of nontranscribed genes: bivalent (marked by both H3K4me3 and H3K27me3 at promoters; red), monovalent (marked by H3K4me3, but not by H3K27me3 at promoters; blue) and nonmarked (marked neither by H3K4me3 nor by H3K27me3 at promoters; black) genes. (C) Average promoter nucleosome organization for three types of genes: with elongating Pol II (red), with nonelongating Pol II (blue), and with H3K4me3 at promoters but without Pol II (black). (A–C) All reads were extended to 147 bp; the middle 73 bp were taken, piled up, and normalized by sequencing coverage. Number of genes used to draw an average profile was indicated for each class. Short genes (<1 kb) were excluded. The accompanying nucleosome profiles for each class of genes in the normalized version and at the 256-cell stage are shown in Supplemental Figure S10.











