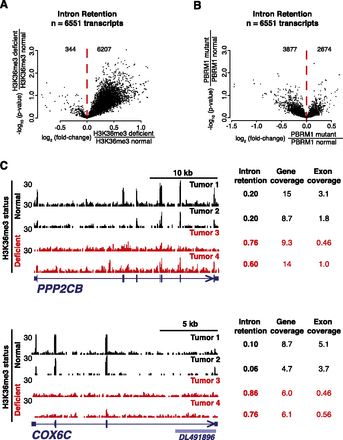
H3K36me3 deficiency is associated with intron retention. Intron retention scores for selected genes (Supplemental Fig. S1C) were compared between (A) H3K36me3-deficient tumors and H3K36me3-normal tumors, and (B) PBRM1-mutant and PBRM1-normal tumors. (C) Example genes exhibiting increased intron retention in H3K36me3-deficient tumors (top, PPP2CB; bottom, COX6C). Intron retention scores, genic coverage (calculated with both intron and exon reads), and exonic coverage (calculated only with exonic reads) are provided for two H3K36me3-deficient tumors (red) and two H3K36me3-normal tumors (black).











