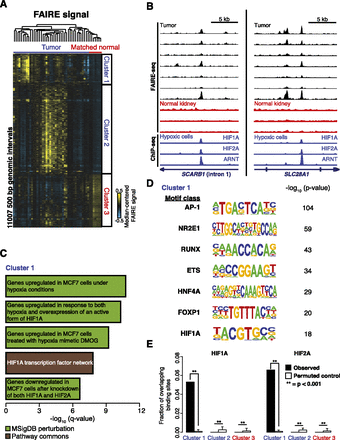
Regions of tumor-specific nucleosome eviction identify the underlying role of HIF in ccRCC. (A) Hierarchical clustering of median-centered FAIRE signal in windows with significant differences between tumors and normal kidney (two-sided t-test, P < 0.01). (B) FAIRE-seq tracks for ccRCC (black) and uninvolved kidney (red) at two loci. (Blue) ChIP-seq signals (Schodel et al. 2011) from HIF1A, HIF2A, and ARNT. (C) The top five Gene Ontology associations (q < 1 × 10−5) with sites in Cluster 1 are shown. (D) Transcription factor binding motifs enriched in Cluster 1 compared with local background 500-bp flanking windows (>2.5-fold over background and present in at least 10% of the Cluster 1 windows). P-values relative to local background are shown. (E) Fraction of HIF1A and HIF2A binding sites (Schodel et al. 2011) that overlap the loci in Clusters 1, 2, and 3 compared with permuted controls. Errors bars represent standard deviation (SD).











