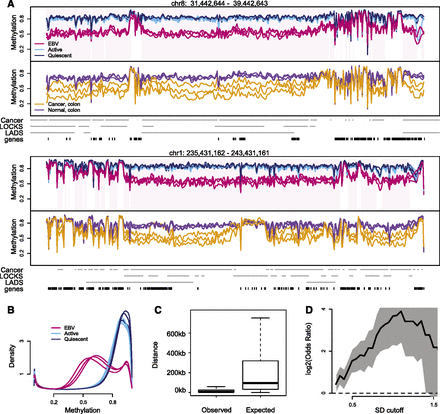
Large hypomethylated genomic blocks in EBV-immortalized B-cells. (A) Smoothed methylation values from bisulfite sequencing data for quiescent (dark blue), activated (light blue), and EBV immortalized (red) B-cells, top panel. The smoothed methylation values estimate average DNA methylation on the kilobase scale. Hypomethylated EBV blocks are demarcated in pink shading. The bottom panel shows smoothed DNA methylation values for normal colon (purple) and colon tumor (orange) samples, from Hansen et al. (2011). (B) Genome-wide distribution of DNA methylation. The large block domains appear as a large bump around 0.6. (C) Simulations show that block locations co-occur. For each of the three EBV transformed samples, we find sample-specific blocks by comparing the sample in question to all three activated samples. For each set of sample-specific blocks, we computed the distance from the observed start position of each sample-specific block to the closest start position in the other two sets. The boxplot on the left shows the distribution of these distances, pooled across all six comparisons. The boxplot on the right shows the expected distribution of distances under the null hypothesis that the block start positions do not agree. The smaller values seen in the left boxplot demonstrates that the start positions of the sample-specific blocks co-occur much more frequently than expected by chance. (D) Enrichment of hypervariable genes in EBV-transformed cell lines, inside EBV blocks. The x-axis denotes a standard deviation cutoff, above which genes are considered hypervariable. The y-axis is the log2 odds ratio of enrichment of these hypervariable genes inside EBV blocks. The gray shaded area is a 95% confidence interval, and values above 0 mark enrichment.











