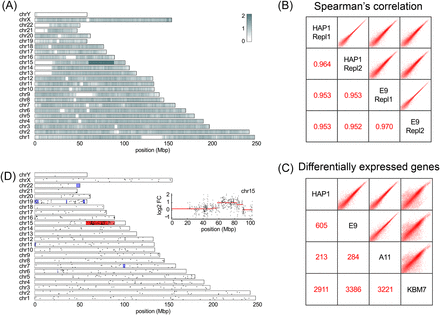
Genomic and transcriptomic changes in eHAP cells are largely confined to Chromosome 15. (A) Whole-genome sequencing was performed on parental HAP1 cells and the A11 and E9 eHAP clones. In this panel, relative coverage between HAP1 and E9 data reveals a copy number loss restricted to the edited Chr 15 fragment. Large white regions correspond to unassembled pieces of the human genome. (B) Two biological replicates of HAP1 cells and two technical replicates of the E9 clone were subjected to RNA sequencing. Spearman’s correlation between the samples shows that overall expression is consistent between the parental line and the edited clones. (C) Two replicates of each cell line were compared pairwise. The number of highly expressed (FPKM > 5) and twofold differentially expressed genes are indicated. (D) Expression ratios between HAP1 and E9 cells were subjected to segmentation analysis. Red/blue bands show segments for which the expression level is consistently higher/lower in the parental cell line. (Inset) Details of the segmentation.











