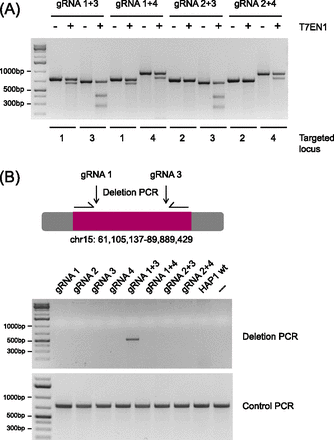
Combination of gRNAs 1 and 3 causes deletion of the Chromosome 15 fragment. HAP1 cells were transfected with various combinations of gRNAs (as indicated). Around 10 d post transfection, genomic DNA was isolated from pools of transfected cells. (A) The regions targeted by individual gRNAs were amplified by PCR using suitable primer pairs. Digestion of these PCR products by T7 endonuclease provides a semiquantitative measure for Cas9 editing efficiency. (B) To assess whether the fragment between gRNAs 1 and 3 had been excised following Cas9 cleavage, we performed a deletion PCR using a forward primer (HG6090) that binds to position Chr 15: 61,105,055 and a reverse primer (HG6093) that binds to position Chr 15: 89,889,818. We also included a control PCR (primer pair HG6090/HG6091) to confirm that every sample contained genomic DNA, suitable for PCR.











