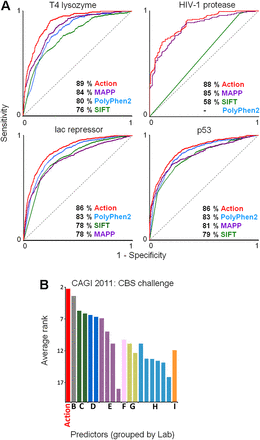
The performance of the Evolutionary Action method was compared to state-of-the-art methods. (A) The area under the receiver operating characteristic curve (AUC) of the relative sensitivity and specificity to separate harmful from harmless mutations for the Evolutionary Action, PolyPhen-2, SIFT, and MAPP was calculated for each of the data sets: 2015 bacteriophage T4 lysozyme mutants to break the host cell walls; 4041 E. coli lac repressor mutants to repress β-galactosidase more than 20-fold; 336 HIV-1 protease mutants to cleave the Gag and Gag-Pol precursor proteins (PolyPhen-2 returned no predictions for the HIV-1 protease mutations); and 2314 human TP53 mutants to transactivate eight TP53 response-elements in yeast. (B) The average rank of current methods (bars), from different groups (letters), to predict the activity of cystathionine beta-synthase (CBS) mutants was assessed by the Critical Assessment of Genome Interpretation (CAGI) of 2011. The CBS activity was assayed for the ability of each mutant to restore growth in yeast cells lacking the normal CYS4 ortholog under two different growth conditions (high and low concentrations of pyridoxine cofactor) (Mayfield et al. 2012). Twenty methods from nine groups were assessed over nine criteria (precision, recall, accuracy, harmonic mean f1, Spearman’s rank correlation coefficient, Student’s t-test P-value, root mean square deviation [RMSD], RMSD over Z-scores, and the AUC) for each cofactor concentration, and then their rank was averaged. Evolutionary Action is shown in red, and a taller bar is a better rank. Raw data and assessment details are available at the CAGI website (https://genomeinterpretation.org/) and from the CAGI organizers Susanna Repo, John Moult, and Steven E. Brenner. The Evolutionary Action analysis files are available at http://mammoth.bcm.tmc.edu/KatsonisLichtargeGR.











