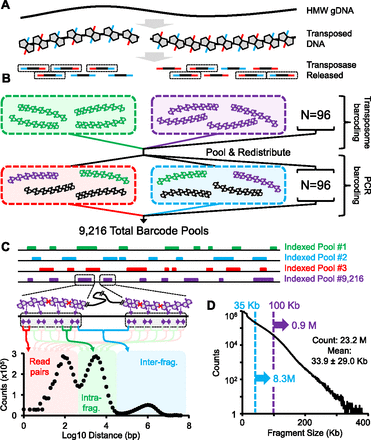
CPT-seq method and performance. (A) High molecular weight (HMW) genomic DNA reacted with hyperactive Tn5 transposase loaded with indexed adaptors. After the transposase complex fragments the DNA and appends the indexed adaptors, the enzyme remains tightly bound to the DNA, such that library molecules derived from the same HMW genomic DNA molecule remain physically linked. Once the transposase is removed by denaturation, PCR amplification of viable templates (gray boxes) can be performed. (B) Schematic of two tier indexing. A 96-plex indexed tagmentation is performed (but without removing the transposase), followed by pooling, mixing, and redistribution to 96 wells. These new pools are subjected to removal of the transposase, 96-plex indexed PCR and then pooling to a single sequencing library. Individual molecules within the final library have indices corresponding to both the pool in which their originating HMW genomic DNA fragment was present during tagmentation (96 indices) as well as during PCR (96 indices), such that there are effectively 96 × 96 = 9216 compartments. (C) Representation of coverage profiles for indexed fragment pools, i.e., compartments (top) and trimodal distribution of adjacently aligning reads within individual compartments. The first peak (∼100 bp; red) corresponds to simple read pairs; the second peak (∼3.2 kbp; green) corresponds to reads originating from the same HMW genomic DNA fragment; the third peak (∼1 Mbp; blue) corresponds to reads originating from different HMW genomic DNA fragments. (D) Distribution of estimated HMW genomic DNA fragment lengths for CPT-seq of GM12878. The mean fragment size is 33.9 kbp, but it is a broad distribution and nearly 1M fragments are >100 kbp.











