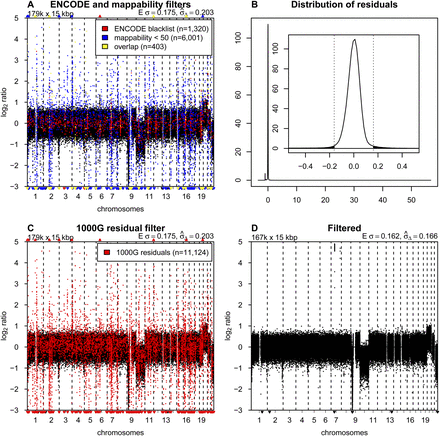
Blacklisting problematic regions. (A) Copy number profile for sample LGG150 with bins overlapping with the ENCODE blacklist highlighted in red, bins with mappabilities below 50 highlighted in blue, and the overlap between the two in yellow. (B) Distribution of median residuals per bin from the 1000 Genomes Project across the 38 samples. Residuals are defined as the distance between observed read counts and the fitted LOESS surface, divided by the LOESS value. The outer plot shows the entire range of values with two discrete peaks. The minor peak around −1.0 results from repetitive sequences. Reads that align equally well to multiple locations in the genome are filtered out. Repetitive sequences therefore have a lower than expected number of reads mapped. The major peak around zero contains most of the bins, and the inset shows a magnification of the peak, with the dotted vertical bars and the shaded area showing the cutoff of 4.0 standard deviations (as estimated with a robust first-order estimator) for blacklisting. (C) Copy number profile of sample LGG150 with bins in the novel blacklist based on residuals of the 1000 Genomes samples highlighted in red. (D) The final copy number profile of sample LGG150 after filtering out bins in the ENCODE and 1000G blacklists.











