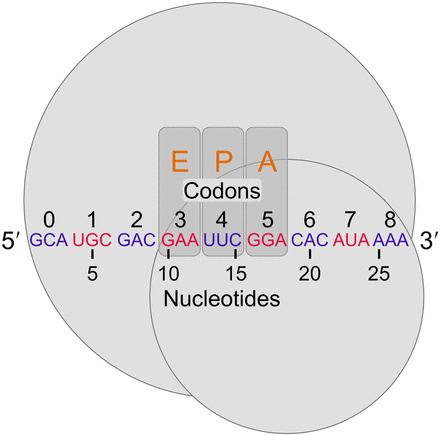
Defining positions relative to the 5′ end of riboprofiling reads. Following the mapping approach of Ingolia (2010), ribosomes (large and small subunits represented by gray circles) protect at least 27 nt of mRNA, corresponding to at least nine codons. Nucleotides and in-frame codons were counted from 5′ to 3′ as shown (arbitrary codons are indicated in alternating blue and red for clarity). In the figure, the ribosome-protected fragment begins in the first reading frame within a codon. However, for reads mapping to the second or third reading frames, while nucleotide counting begins at the first nucleotide, codon counting remains in-frame with the first codon, 0, corresponding to the one containing the first nucleotide. For reference, the orange letters indicate the codons that previous studies have indicated as the exit-tRNA (E-site), the peptidyl-tRNA (P-site), and aminoacyl-tRNA (A-site) sites, respectively (Ingolia et al. 2009; Stadler and Fire 2011; Li et al. 2012; Qian et al. 2012; Zinshteyn and Gilbert 2013).











