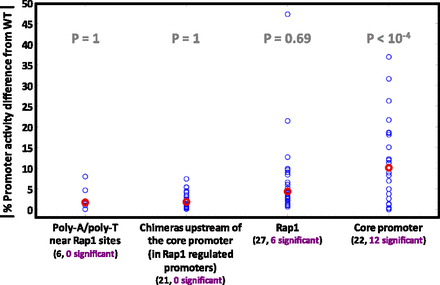
The effect of various types of orthologous mutations on promoter activity. Most orthologous mutations of RP promoters fall into four categories: poly(dA)/poly(dT) tracts adjacent to Rap1 binding sites; chimeras of Rap1-regulated RP promoters, where entire promoter prefixes upstream of the core promoter region were replaced between orthologs; Rap1 binding sites (and possibly their adjacent context); and core promoter mutations. For each mutation, its absolute percent difference in promoter activity from the wild type is shown by a blue circle. Red circles show median values. A chimera was regarded to have a significant promoter activity difference only if its promoter activity was significantly different from those of both native RP promoters. Below each category name appears the total number of mutations (black font) and the number of those with promoter activity significantly different from the wild type (purple font). For each category, a significant mutation enrichment P-value (one-sided hyper-geometric test) is in gray.











