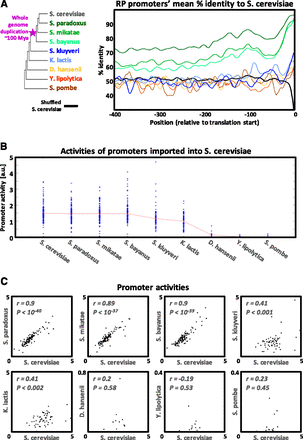
Promoter sequence and activity divergence for orthologous yeast RP promoters. (A) The mean orthologous sequence identity to S. cerevisiae along RP promoters of different yeast species, computed based on pairwise alignments (smoothed using a 21-bp sliding window). The color of each track corresponds to the color of the yeast species name appearing in the left phylogenetic tree. The black track is the result of aligning S. cerevisiae promoters to their own randomly shuffled sequence. The big dip in the black track toward its downstream edge is due to the extremely high A content of the S. cerevisiae RP promoters just upstream of the ORF (see Supplemental Fig. 2). (B) Measured native RP promoter activities for each species. Median values for each species appear in red. (C) Each dot plot compares the measured activities of RP promoters from one species versus their S. cerevisiae orthologs.











