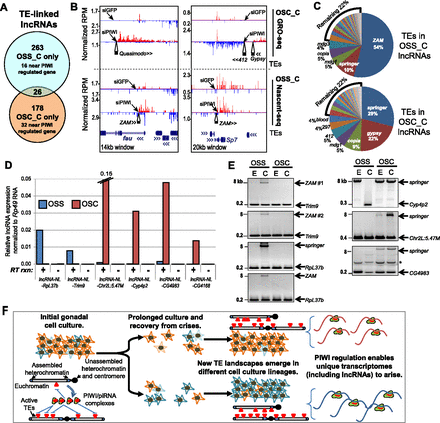
TE landscape differences correlate with lncRNA diversity. (A) Venn diagram of unambiguous lncRNAs that are mostly distinct populations between OSCs and OSS cells. The overlap represents lncRNAs that overlap by at least 1 kb in genomic coordinates from the D. melanogaster Release 5/dm3 genome. (B) Representative loci where both OSC and OSS cell genomes contain TE insertions, but the differences in TE composition are linked to differences in lncRNA configurations. (C) Composition of the major TE classes falling within lncRNA loci. (D) Quantitative RT-PCR of lncRNA expression, normalized to Rp49 levels, for loci where TE insertions are present in just one follicle cell line (lncRNA-NL-Trim9 and -NL-RpL37b in OSS cells; -NL-Chr2L:5.47M, -NL-Cyp4p2, -NL-CG4983, and -NL-CG4168 in OSCs). (E) Genomic PCR confirmation of de novo TE insertions that are either only in OSCs or only in OSS cells. Asterisk marks a nonspecific amplicon, and sequence-specific obstacles in primer design prevented genomic PCR validation of TE insertions in the CG4168 locus. (F) Model for TE dynamics in follicle cell cultures that result in distinct TE landscapes and unique transcriptome profiles, including the production of novel lncRNAs.











