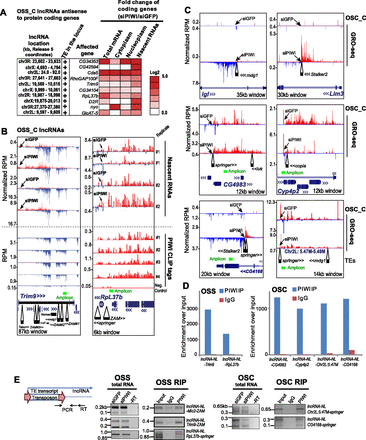
Novel long noncoding RNAs (lncRNAs) are stimulated by de novo TE insertions. (A) Heat map of representative OSS cell genes overlapped by lncRNAs and up-regulated during PIWI knockdown. D. melanogaster Release 5/dm3 reference genome coordinates are shown here; updated Release 6 coordinates are in Supplemental Figure S8E. (B) Representative lncRNA loci in OSS cells, with nascent RNAs in upper tracks and PIWI CLIP tag profiles in lower tracks. The lncRNA-NL-Trim9 is on the minus strand (blue reads), while the lncRNA-NL-RpL37b is on the plus strand (red reads). Arrows point to coding transcripts for Trim9 (red reads) and RpL37b (blue reads) that increase when PIWI is knocked down. Location of de novo TE insertions are at the bottom of the diagrams, whereas green bars mark amplicons for evaluating lncRNA enrichment in a PIWI RIP experiment and RT-qPCR. (C) Representative TE-associated lncRNAs in OSC cells identified from GRO-seq reads. The lncRNAs at igl and Lim3 are unambiguously antisense to the coding transcripts, whereas the transcripts overlapping the same sense of CG4983 and Cyp4p2 may not adhere to the strict definition of lncRNAs, but their long extensions in noncoding regions are reminiscent of defined lncRNAs. Arrows point to the putative start of the lncRNA. (D) RIP assay validates lncRNA association with PIWI. The lncRNAs at igl and Lim3 were not analyzed by PIWI RIP, since they are expressed only upon PIWI knockdown. (E) RT-PCR of amplicons that span the inserted TE sequence and the lncRNA. These data are consistent with lncRNA nascent reads not affected by PIWI knockdown and enrichment of lncRNA in PIWI RIP.











