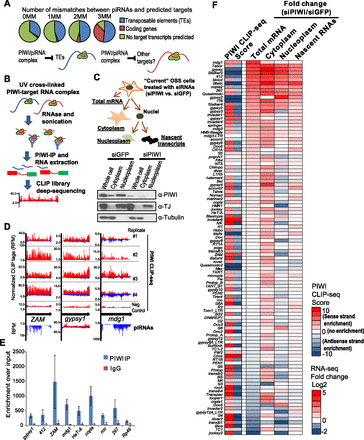
Transcriptome profiling and CLIP-seq confirm that transposable elements (TEs) are the main direct targets of PIWI-mediated regulation. (A) Bioinformatic prediction of PIWI/piRNA targets in OSS cells based on complementarity to piRNAs (Lau et al. 2009). With more mismatches (#MM), more coding genes are predicted to pair with piRNAs. These data suggest possible PIWI targets beyond TEs. (B) Scheme for PIWI CLIP-seq approach to identify PIWI associated transcripts. (C) Scheme for cellular fractionation and mRNA and nascent RNA isolation from OSS cells treated with siRNAs. Western blots confirm successful PIWI knockdown and separation of cytoplasmic from nucleoplasmic fractions. TJ is a transcription factor marking nuclear fractions, while tubulin is mainly cytoplasmic. (D) Different PIWI CLIP-seq profiles and corresponding piRNAs profiles of TEs with top PIWI CLIP scores. (Red) Plus strand reads; (blue) minus strand reads. Normalized CLIP-seq reads were deemed significant from our CLIP-seq processing algorithm. (RPM) Reads per million. (E) RNP-IP (RIP) assays validate TE transcript association with PIWI. Error bars correspond to standard deviation from five biological replicates. (F) Heat map showing TE transcript level changes in different compartments after PIWI knockdown in comparison to PIWI CLIP-seq scores. PIWI CLIP-seq scores with only red colors reflect primarily sense-strand patterns (Fig. 1D), only blue colors reflect primarily antisense-strand patterns (Supplemental Fig. S2F), and both strands when both red and blue colors are shown.











