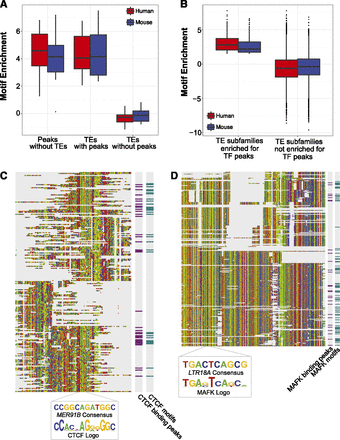
Binding motifs of TFs were enriched in TE-derived binding peaks. (A) Distribution of motif enrichment scores (log-odds ratio) (see Methods) in non-TE, TF binding peaks (training set for de novo motif prediction), TE-derived TF binding peaks (test set), and TEs without peaks (control). (B) Distribution of motif enrichment scores in TE subfamilies that were enriched in TF binding peaks compared with all other TE subfamilies. (C) Multiple sequence alignment of MER91B genomic copies (n = 215) from the mouse genomes and the MER91B consensus sequence (bottom row of the alignment). Beside the alignment are indications of genomic copies that had CTCF binding peaks (purple) and CTCF motifs (green). (D) Similarly, multiple sequence alignment of LTR18A genomic copies (n = 259) from the human genome along with the LTR18A consensus sequence (bottom row of the multiple-sequence alignment). Annotated on the right are indications of genomic copies that had MAFK binding peaks and MAFK motifs. Nucleotides in alignments are color-coded: (A) green; (C) blue; (G) yellow; (T) red.











