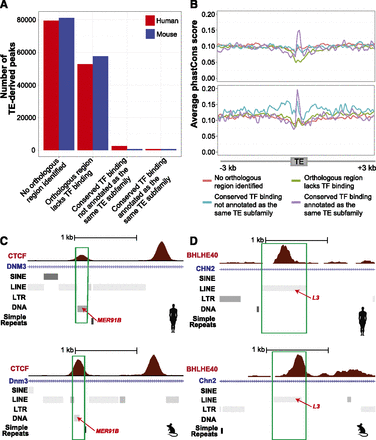
The majority of TE-derived binding peaks were species-specific, whereas the shared ones exhibited sequence conservation. (A) Bar plot showing the numbers of binding events in various categories of conservation. We classified TE-derived binding events in each species into four categories: (1) no orthologous region identifiable for the TF binding peak; (2) orthologous region lacks TF binding; (3) conserved TF binding, not annotated as the same TE subfamily in both species; and (4) conserved TF binding, annotated as the same TE subfamily in both species. (B) Distribution of phastCons scores (see Methods) in 6-kb windows centered on the TE sequences in the four categories defined above. The signal was averaged across 25-bp bins and smoothened for this plot. TE-derived binding peaks that could be mapped to binding peaks derived from the same TE subfamily in the other species exhibited increased sequence constraint over the background. (C,D) UCSC Genome Browser images of TE-derived TF binding peaks, whose occupancy was conserved between human (upper panel) and mouse (lower panel). The browser images correspond to (C) CTCF binding encoded on MER91B fragments in human and mouse, and (D) BHLHE40 binding encoded on L3 fragments in human and mouse.











