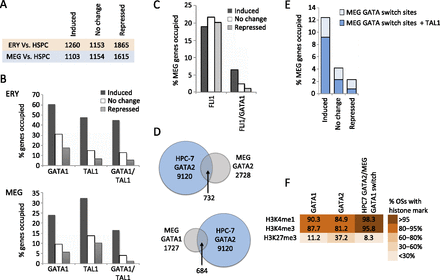
Functional annotation of GATA1, GATA2, and FLI1 occupancy in megakaryopoiesis. (A) Numbers of up-regulated (induced), unchanged, and down-regulated (repressed) genes in ERY and MEG compared to HSPCs, used as a reference point. (B) Fractions of up-regulated, unchanged, and down-regulated genes (versus HSPC) occupied by GATA1 or TAL1 (irrespective of co-occupancy by other transcription factors) and genes co-occupied by GATA1 and TAL1 in ERY and MEG. (C) Fraction of up-regulated, unchanged, and down-regulated genes occupied by FLI1 in MEG (all FLI1 OSs and those co-occupied by GATA1 and FLI1). (D) Intersection of transcription factor occupancy peaks between MEG and the multipotent hematopoietic cell line HPC-7 (Wilson et al. 2010). (E) Fractions of up-regulated, unchanged, and down-regulated genes containing a GATA switch site (replacement of GATA2 with GATA1) in MEG development, with or without concurrent TAL1 binding. (F) Color-coded fractions of transcription factor binding sites associated with H3K4me3, H3K4me1, and H3K27me3 histone methylation patterns in MEG.











