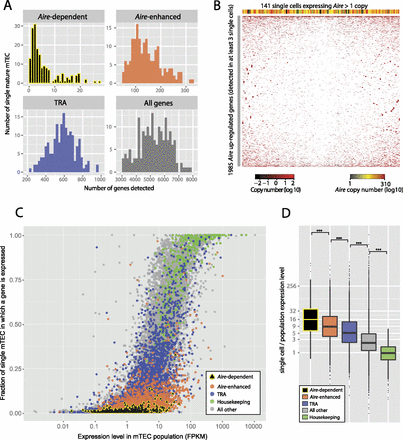
Transcriptomic analysis of promiscuous gene expression in single TEC. (A) Single mature mTEC tend to express few genes that are dependent on or enhanced by Aire expression (as defined in Fig. 3C). The histograms show the number of genes detected in 174 single mature mTEC that expressed >3000 protein-coding genes. (B) No discernible clustering is evident from the hierarchical clustering (with optimized leaf ordering) of 141 single Aire-expressing mature mTEC (columns) and 1985 genes up-regulated by Aire expression (rows) detected in at least three of these single cells. The colored bar above the plot indicates the single-cell expression level of Aire. (C) Genes dependent on Aire expression are transcribed less frequently in single mature mTEC than are other genes. The scatter plot shows the fraction of single mature mTEC that express any given gene against the expression level of that gene determined from the mature mTEC population. (D) When transcribed in single mTEC, genes dependent on Aire expression tend to be present at a level 16-fold higher than that indicated by the population average. Before calculating the relative expression levels, single-cell gene expression levels were globally normalized against population values using a linear model. (***) P < 1 × 10−14, estimated using the Mann-Whitney U-test.











