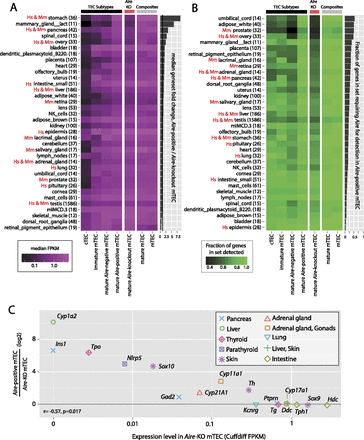
Requirement for Aire reflects known AIRE deficiency pathologies. (A) The median expression level (FPKM) of sets of genes restricted in expression to single physiological samples (excluding the thymus) of the GNF GeneAtlas (based on dynamic step criteria; see Methods; Supplemental Fig. 6) (gene numbers indicated in parentheses) for each TEC population. Gene sets are sorted by the fold change in median expression level between mature Aire-positive and mature Aire-KO mTEC (accompanying bar chart). In both A and B, gene sets representing organs affected by AIRE deficiency in APS-1 (“Hs”) and the corresponding mouse model (“Mm”) are indicated. (B) The fraction of the same sets of genes that are detectable (<5% local FDR) in each TEC population (see Methods; Supplemental Fig. 3); gene sets are sorted by the increase in the fraction of these genes detected in mature Aire-positive mTEC compared to Aire-KO TEC. (C) Relative expression of known APS-1 autoantigens in mature Aire-positive wild-type and knockout TEC (Shikama et al. 2009). Induction by Aire expression is significantly negatively correlated with the transcriptional level of these genes in mature Aire-KO mTEC.











