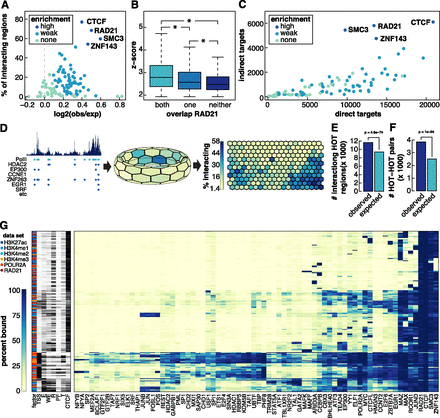
Factors enriched at interacting loci. (A) TF enrichment at interacting loci. The x-axis represents the log2 ratio of observed divided by expected TF binding peaks that overlap interacting loci. The y-axis represents the number of interacting loci at which that factor is bound. Colors of circles represent the level of enrichment (see Supplemental Methods). (B) Box and whisker plot of Z-scores of interactions that overlap a RAD21 peak at both, one, or neither end of an interaction. (*) Significant differences (P < 0.05, Wilcoxon signed-rank test). (C) Scatter plot comparing the number of direct and indirect targets of each TF. Colors correspond to how enriched that factor was at interacting loci. (D) Depiction of how SOM maps were generated using POLR2A as an example. (Left) Binding profile of POLR2A in K562 cells as well as POLR2A peaks (light blue circles) and peaks of other TFs that overlap POLR2A peaks (dark blue). (Middle) Toroidal depiction of SOM generated from POLR2A data set. Each hexagon represents a neuron comprised of POLR2A binding peaks that share patterns of TF cobinding. (Right) Planar view of POLR2A SOM map. Neurons are colored to depict the percentage of POLR2A peaks in each neuron that are involved in an interaction. (E) Barplot showing number of observed and expected HOT regions involved in interactions. P-values were determined by Fisher’s exact test. (F) Barplot showing number of observed and expected number of interactions linking two HOT regions. Significance was determined by permutation testing. (G) Heatmap representing neurons that are significantly enriched for interactions (P < 0.01, Fisher’s exact test with Benjamini-Hochberg correction, fold-enrichment > 2). For visualization purposes, only selected TFs are shown. The data sets in which a neuron was detected are shown on the far left as well as the percentage of peaks in each neuron that overlapped different types of annotated DHSs.











