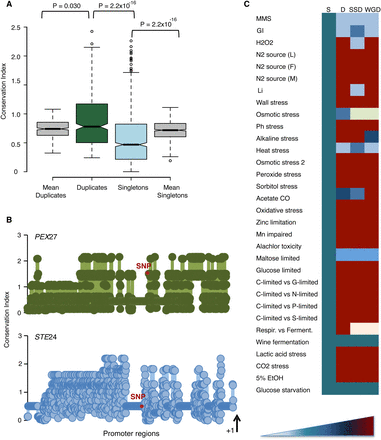
Duplicates show more expression plasticity than singletons under stress conditions. (A) Mutations at duplicate promoters occur at more conserved regions than those at singleton promoters. Conservation coefficient
is calculated by measuring the amount of entropy in each nucleotide site of the alignment that comprised upstream regions
of genes in S. cerevisiae and at least five other closely related orthologs. Duplicates show larger conservation in their mutated sites than expected,
while singletons show less conservation than expected. (B) Two examples of the conservation of mutated sites at duplicate (PEX27) and singleton (STE24) promoter regions. Red dots represent mutated nucleotide sites during our evolution experiment. The first site from the initiation
codon is also labeled (+1). (C) We analyzed 32 stress conditions from various independent studies (Supplemental Table S7). The phenotypic (expression) plasticity
of genes, both the duplicates (D) and singletons (S), was calculated as the difference in the expression of the gene between
two environmental conditions (Ei and Ej): 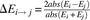 . Duplicates with larger expression plasticity are colored in red; squares are colored in blue that becomes lighter as the
difference in expression decreases between the duplicates; and light yellow indicates that the corresponding information is
not sufficiently large to perform statistical tests.
. Duplicates with larger expression plasticity are colored in red; squares are colored in blue that becomes lighter as the
difference in expression decreases between the duplicates; and light yellow indicates that the corresponding information is
not sufficiently large to perform statistical tests.











