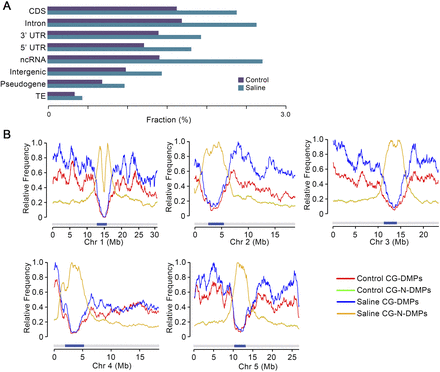
Genome-wide distribution of CG-DMPs in saline soil and control MA lineages. (A) Distribution of CG-DMPs in genomic regional categories as determined by genome sequence annotation. (CDS) Coding DNA sequence; (UTR) untranslated region; (ncRNA) noncoding RNA; (TE) transposable element, expressed as fraction of total methylated GC sites in all saline soil and control G10 samples. (B) Distribution along individual chromosomes (Chr 1, Chr 2, etc.) of CG-DMPs and CG-N-DMPs (see text for definition of CG-N-DMPs). Data from G10 saline soil and control samples were normalized to the highest value for each chromosome and class (CG-DMPs or CG-N-DMPs). Dark blue bars indicate centromeres. Because Control and Saline CG-N-DMPs track very closely, the latter trace substantially obscures the former.











