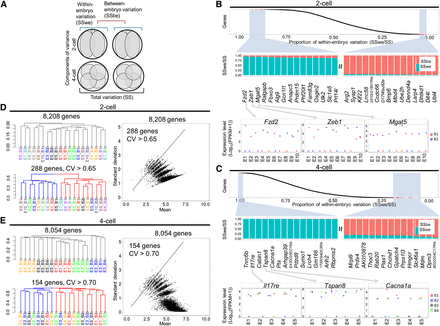
Variance decomposition and unsupervised clustering of single cells. (A) The total variation (SS) can be decomposed into the sum of within-embryo variation (SSwe) and between-embryo variation (SSbe). (B,C) Genes (y-axis) are rank-ordered from the largest to the smallest by SSwe/SS (x-axis). Expanded views of the genes with the largest and the smallest SSwe/SS are shown beneath, followed by the expression levels of selected genes. (E1–E10) Embryos 1 to 10. (B1, B2) Blastomeres 1 and 2. (FPKM) Upper quartile normalized FPKM. (D,E, left) Clustering of the blastomeres using all mRNAs (upper dendrogram) and high CV genes (lower dendrogram). (Ei-Bj) The jth blastomere of embryo i. (Right) A scatter plot of standard deviation (y-axis) vs. mean (x-axis) of all genes. The CV filter selected for the genes above the line.











