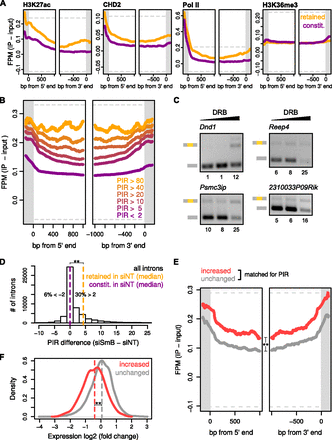
Coupling between IR and RNA Pol II elongation. (A) Average ChIP-seq signals across introns that are retained (orange, PIR ≥ 10) or not retained (purple, PIR < 2) in K562 cells, for four of 129 analyzed human chromatin features. Retained and not retained introns analyzed are in genes with matched expression levels. (Gray bars) Flanking exons; (y-axes) input-subtracted average ChIP fragment density per million reads (FPM-input) at each aligned bp; (gray dashed lines) 2nd and 98th percentiles of all values in the plot region before averaging to give an indication of the dynamic range. See Supplemental Figure S9A for similar analysis in mouse CH12 cells. (B) Average ChIP-seq signal of Ser2-phosphorylated RNA Pol II over introns with different PIR thresholds in K562 cells. Labels and plots as in A. See Supplemental Figure S9B for similar analysis in mouse CH12 cells. (C) RT-PCR assays of IR events in CGR8 mouse ES cells with and without treatment with the RNA Pol II elongation inhibitor DRB. Retained introns with low PIR in ES cells relative to other cell types were analyzed. (Lane 1) 0 μg/mL DRB; (lane 2) 10 μg/mL; (lane 3) 25 μg/mL. Quantitations are shown beneath each panel. See Supplemental Figure S10 for the full set of tested introns. (D) Number of introns with PIR changes in HeLa cells transfected with siRNAs targeting SNRPB, which codes for SmB/B′, relative to cells transfected with a control, nontargeting siRNA (Saltzman et al. 2011). (Dashed lines) Medians of retained (PIR ≥ 10) and constitutive (PIR < 2) introns in control treated cells; (asterisks) P < 0.001 between these groups (one-sided Mann-Whitney U test). (E) RNA Pol II occupancy over introns (in untreated HeLa cells) that become more retained (PIR difference > 15, red) or that do not change (PIR difference between −2 and +2, gray) after SNRPB knockdown. Because RNA Pol II occupancy is dependent on absolute PIR, groups of introns with matching PIR distributions in control treated cells and >2 kb away from the transcription start site are shown. (Asterisks) P < 0.001 for difference in median Pol II-Ser2p occupancy (see Methods for test details). (F) Expression change between SNRPB and control knockdown for genes containing introns that become more retained (PIR difference > 15, red) or that do not change (PIR difference between −2 and +2, gray). (Asterisks) P < 0.001 between these groups (one-sided Mann-Whitney U test).











