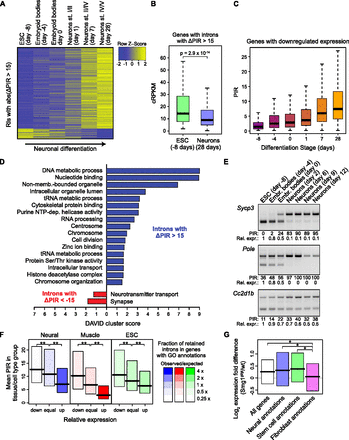
IR-mediated tuning of gene expression. (A) Heatmap of Z-scores for introns differentially retained during differentiation of ES cells into mature glutamatergic neurons. Z-scores are shown for retained introns with a change in PIR (ΔPIR) > 15 between ES cells and mature neurons (day 28). (B) Distribution of expression values (in cRPKM) in ES cells and mature neurons for genes that contain introns with increased retention during differentiation. (C) Distributions of PIR of introns in genes whose expression is down-regulated (more than fivefold between day −8 and day 28) at different time points of neuronal differentiation. (D) DAVID cluster analysis of enriched GO annotations for genes that contain introns with increased (blue) or decreased (red) retention during neuronal differentiation. (E) RT-PCR validation of RNA-seq–detected events with increasing PIR and decreasing spliced mRNA expression during differentiation of glutamatergic neurons from mouse ES cells. Quantification of PIR and the mRNA expression are shown beneath each panel. For relative expression, the spliced band was quantified and normalized to Gapdh, and day −8 was set to 1. See Supplemental Figure S8 for the full set of 11 tested events. (F) Relationship between the degree of IR and cell type-specific expression of genes annotated with cell/tissue-specific functions. Bars show the upper and lower quartile, and black lines the medians, of the PIR difference between neural and other cell/tissue types (left), muscle and other cell/tissue types (middle), and ESCs and other cell/tissue types (right). Intron PIR was measured in sets of genes assigned to equal-sized bins (three bars) based on having decreased (down), equivalent (equal), or increased (up) expression, as compared to the median expression values for the other cell types. Shading indicates the degree of enrichment of GO categories related to the biology of neural, muscle, and stem cells, respectively (see Methods for details). Asterisks indicate P < 0.001 in one-sided Mann-Whitney U tests after Bonferroni correction. (G) Expression difference between Smg1gt/gt and wild-type MEFs for transcripts harboring retained introns predicted to introduce a PTC that can trigger NMD, in genes associated with GO categories related to the biology of neural cells, stem cells, and fibroblasts. Asterisks indicate a significant difference between median values for expression level differences (P < 0.05 in one-sided Mann-Whitney U tests after Bonferroni correction).











