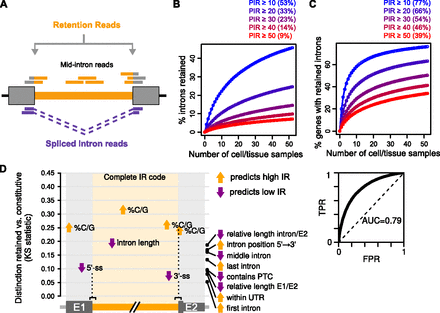
Detection and prevalence of intron retention. (A) Intron retention was detected by aligning RNA-seq reads to comprehensive sets of exon–intron and exon–exon junctions. Reads mapping to mid-intron sequences and balanced counts of reads aligning to upstream and downstream exon–intron sequences were used to discriminate IR from other forms of transcript variation. IR levels were measured using percent intron retention (PIR): 100× mean retention reads over the sum of retention and spliced intron reads (see Methods for details). (B) Percentage of total human introns detected as retained at different PIR thresholds as increasing numbers of cell and tissue samples are randomly sampled. Numbers in parentheses are estimates for percentages of total introns retained at different PIR thresholds, as derived from extrapolation. (Circles) Means from 1,000 iterations; (lines) fitted function used for extrapolation (see Methods). Data for mouse introns in Supplemental Figure S1C. (C) Percentage of total human genes with retained introns at different PIR thresholds as increasing numbers of cell and tissue samples are randomly sampled. Numbers in parentheses are estimates for percentages of total genes with retained introns at different PIR thresholds, as derived from extrapolation. Circles and lines as in B; data for mouse genes in Supplemental Figure S1D. (D) Cis-acting features predictive of IR in human. The main graph quantifies, using the Kolmogorov-Smirnov statistic, how well individual features or a logistic regression model comprising 136 features (“Complete IR code”) (Supplemental Table S2) distinguish retained (PIR ≥ 10) from constitutive (PIR < 2) introns in neural tissues. See Methods for details. The graph on the upper right shows the receiver operating characteristic of the complete IR code with area under the curve (AUC) indicated. (TPR) True positive rate; (FPR) false positive rate. E1/E2 are 5′/3′ exons, respectively.











