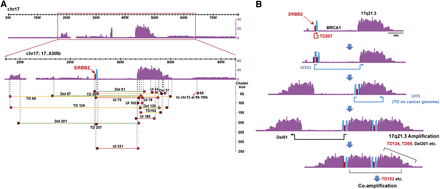
Reconstruction of Chr 17q rearrangements in BT55. (A) Copy number and structural variations mapping to Chr 17 (17–63 Mb) in the BT55 sample. For each structural variation, the brown and pink arrowheads correspond to the left and right mate-pair read clusters, respectively. SV breakpoints map to the end of the brown arrowhead and to the start of the pink arrowhead (as detailed in Hillmer et al. 2011) and are indicated by dashed vertical lines. (TD) Tandem duplication (yellow); (UI) unpaired inversion (red); (Del) deletion (green). The number associated with each structural variation corresponds to its cluster size. (B) A schematic description of the plausible structural events that contributed to the final shape of Chr 17 in the BT55 cancer genome. An initial tandem duplication (TD207) involving the ERBB2 locus was followed by unpaired inversions UI331 and UI75, which is in effect a TD on the cancer genome, resulting in the joining of ERBB2 and the 17q21.3 locus. Subsequent events caused the loss of the BRCA1 locus and the massive coamplification of the ERBB2 and 17q21.3 loci.











