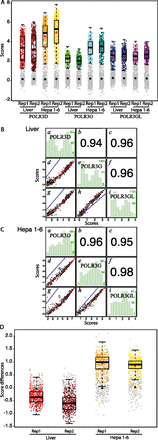
POLR3G and POLR3GL occupy largely the same loci. (A) Box plots showing scores in replicate 1 (Rep1) and replicate 2 (Rep2) samples from liver or Hepa 1-6 cells, as indicated on the x-axis. The y-axis shows scores in log2. Genes with scores below the cutoff (see Methods) are represented by gray dots. The median is indicated by the black horizontal bar, the mean of genes above the cutoff by the red dot, and the mean of genes below the cutoff by the black dot. The four box plots on the left are reproduced from Figure 3C, upper left panel. (B) Spearman's rank correlation of POLR3D and POLR3G (d), POLR3D and POLR3GL (g), or POLR3G and POLR3GL (h) scores in liver cells: In blue, the x = y line; in red, the regression line. (b,c,f) The correlation coefficients corresponding to panels d, g, and h, respectively. (a) Distribution histogram representing, for each POLR3D score interval of 0.5 (see x-axis at the bottom of g), the number of genes in that interval (y-axis at the right of the panel: The numbers in green correspond to the lowest, middle, and highest number of genes). (e) As in a but for each POLR3G score interval of 0.5 (see x-axis at the bottom of h). (i) As in a but for each POLR3GL score interval of 075. (C) As in B but in Hepa 1-6 cells. (D) Box plots showing POLR3G-POLR3GL score differences (in log2) in replicate 1 (Rep1) and replicate 2 (Rep2) samples from liver or Hepa 1-6 cells, as indicated on the x-axis. The y-axis shows score differences (in log2). Genes with scores below the cutoff (see Methods) are represented by gray dots. The median is indicated by the black horizontal bar, the mean of genes above the cutoff by the red dot, and the mean of genes below the cutoff by the black dot.











