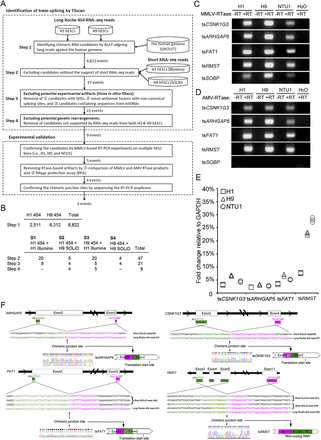
Identification and experimental validation of trans-splicing events in the transcriptome of hESCs. (A) TSscan identification and subsequent experimental validation. TSscan identification involved four steps. The first three steps identified trans-spliced RNA candidates and removed potential in vitro artifacts, and the last step removed potential genetic rearrangements. (SHS) Short homologous sequences. (B) The number of candidates remaining after each TSscan filter step. Note that one candidate may simultaneously belong to different data sets. For example, in Step 2, one candidate belongs to both S1 and S2, and one candidate belongs to both S3 and S4. (C) MMLV-RTase-based and (D) AMV-RTase-based RT-PCR products of tsCSNK1G3, tsARHGAP5, tsFAT1, tsRMST, and tsSOBP in three types of hESC line (H1, H9, and NTU1). (±RT) RT-PCR without/with RTase. (E) qRT-PCR analysis of tsCSNK1G3, tsARHGAP5, tsFAT1, and tsRMST in multiple hESC lines (H1, H9, and NTU1). (F) Schematic representations (top) and sequence chromatograms (bottom) for tsCSNK1G3, tsARHGAP5, tsFAT1, and tsRMST. The long/short RNA-seq reads that support the chimeric junction sites (indicated by arrows) of the corresponding trans-spliced RNAs are shown.











