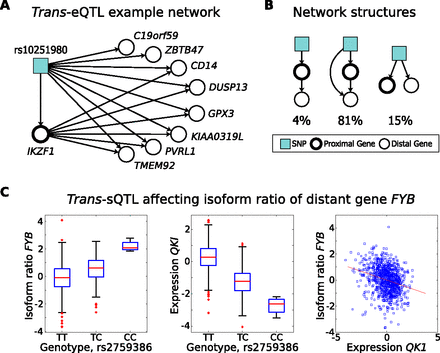
Trans-regulatory variation and mediation through proximal genes. (A) Example subnetwork of significant associations centered on expression of IKZF1. The SNP rs10251980 is associated with expression of the nearby gene IKZF1 (P < 10−8), along with eight distant genes (P < 10−12). IZKF1 is also coexpressed with six of the eight genes (P < 0.05). (B) Prevalence of candidate regulatory network structures for trans-eQTLs including the SNP, the corresponding distant genes, and any genes proximal to the SNP that are also associated with its genotype. For each trans-eQTL gene, we analyzed its relationship with the most strongly associated SNP, along with all genes within 1 Mb of that SNP. Network structures best fitting each set were identified using likelihood ratio tests (Methods; Supplemental Material). (C) Association between rs2759386 and isoform ratio of FYB, potentially mediated through expression of splicing factor QKI, which is proximal to the SNP. Rs2759386 is associated with total expression levels of QKI, and both this SNP and QKI are associated with isoform ratio of FYB (P < 10−14 and P < 10−16, respectively).











