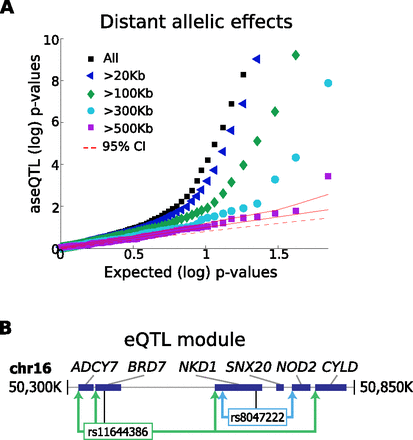
Distant and modular intra-chromosomal regulation. (A) Q-Q plot of aseQTL P-values for intra-chromosomal eQTLs of varying distances. For eQTLs implicating SNPs beyond each distance threshold from the corresponding TSS (0 kb, 20 kb, 100 kb, 300 Kb, 500 kb), we computed aseQTL association tests between the eQTL SNP and allelic ratios at all exonic loci available for the corresponding gene, taking the best association identified from these. The expected P-value distribution and 95% CIs were computed empirically from repeated random draws of SNPs similarly tested against exonic loci within each eQTL gene. We observe that distant eQTLs show more ASE than expected by chance, although the enrichment declines with distance. (B) Schematic of a genomic region on chromosome 16 containing coregulated genes, along with nearby genes and SNPs having an impact on each gene. Rs11644386 affects a discontinuous group of genes, with the farthest association (CYLD) being >400 kb away, and does not have significant associations with two intermediate genes SNX20 and NOD2. Another SNP, rs8047222, is associated with expression of NKD1 and a nearby gene NOD2, but has no influence on the more distant genes BRD7 and ADCY7.











