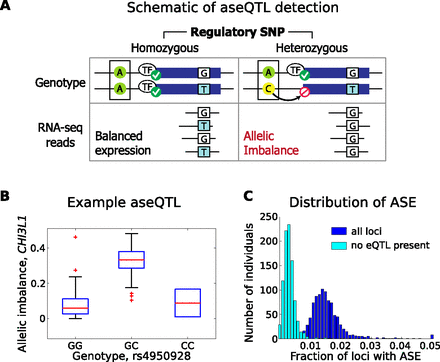
Cis-regulatory variation and allelic effects. (A) Schematic illustration of aseQTL. Heterozygosity at a regulatory locus is linked to allelic imbalance detected from RNA-seq reads over a separate heterozygous coding SNP (a second, separate locus) in the corresponding gene. Conversely, individuals who are homozygous at the regulatory SNP will show balanced allelic expression at the coding SNP (still estimated among individuals who are heterozygous at the coding SNP). (B) Example of aseQTL, the most significant association in this analysis. Rs4950928, a known asthma risk variant SNP in the 5′ UTR of CHI3L1, is associated with allelic imbalance in the coding region of CHI3L1, with heterozygous individuals showing significantly increased allelic imbalance compared to individuals homozygous for either the reference or nonreference allele (P < 10−71) (Methods). (C) Distribution of significant ASE by individual. In each individual, we evaluate the fraction of testable heterozygous loci (requiring sufficient read depth and other filters) (see Supplemental Material) with significant ASE (binomial P ≤ 10−3). To evaluate the distribution of ASE not explained by heterozygosity for a common regulatory variant, we then evaluate the same set of testable loci, but only counting ASE when the individual is not heterozygous for a corresponding eQTL SNP. In this case, we consider SNPs that are significant at P ≤ 10−3 for the corresponding eQTL gene.











