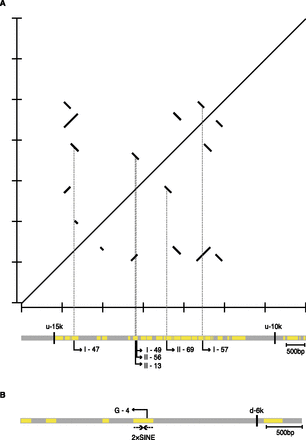
Clustering of DEL breakpoints in genomic inverted repeats. (A) Centromeric DEL breakpoints (rightward bent arrows) in six out of nine 129-DEL or B6-DEL clones (Fig. 3) are located within a distinct 4.2-kb repeat-rich region between SNP markers u-15k and u-10k (Supplemental Table S1). Yellow rectangles represent repetitive sequences. A self-alignment dot plot using BLAST2 (Tatusova and Madden 1999) of a DNA fragment (chr8: 125082686–125090184) containing the 4.2-kb repeat-rich region is shown. Lines with a slope of +1 indicate direct repeats, while lines with a slope of –1 indicate inverted repeats. (B) The telomeric DEL breakpoint (leftward bent arrow) in clone G-4 is localized within tandemly inverted (i.e., perfectly palindromic) 2 × 135-bp SINE repeats. (d-6k) SNP marker located 6 kb telomeric to the Aprt locus (Supplemental Table S1).











